PHARE
Physiology of Aquatic species: REgulations from genes to organisms

Team Members
- Clothilde BERTHELIN, Assistant Professor, MERSEA, PHAREclothilde.berthelin@-Code to remove to avoid SPAM-unicaen.fr, +33 2 31 56 51 14
- Natacha CLAIRET, Temporary Teaching and Research Attaché, MERSEA, PHAREnatacha.clairet@-Code to remove to avoid SPAM-unicaen.fr
- Nathan MORANDI, Doctoral Student, , MERSEA, PHAREnathan.morandi@-Code to remove to avoid SPAM-etu.unicaen.fr
Research team overview
The team brings together University (assistant-) Professors who specialize in the molecular regulation of physiological processes in aquatic species, exploring levels of observation ranging from genes to cells, tissues, and whole organisms.
These regulatory mechanisms are studied both at the intrinsic level (e.g., gene expression control in cis and trans, neuroendocrine communication, cell signaling) and at the extrinsic level (e.g., environmental influences, acclimation to external stimuli). Research questions focus on understanding the mechanisms of cell fate determination and plasticity (phenotypic identity, differentiation) that govern general physiology and biological cycles, particularly in relation to reproduction (sex determination, gametogenesis, spawning) and development (ontogenesis), as well as the stability of these processes in the context of global environmental change.
The team develops its research with an integrative perspective, considering interactions across scales, between physiological systems (e.g., reproduction/development – nutrition – immunity), and between intrinsic and extrinsic factors.
Biological Models
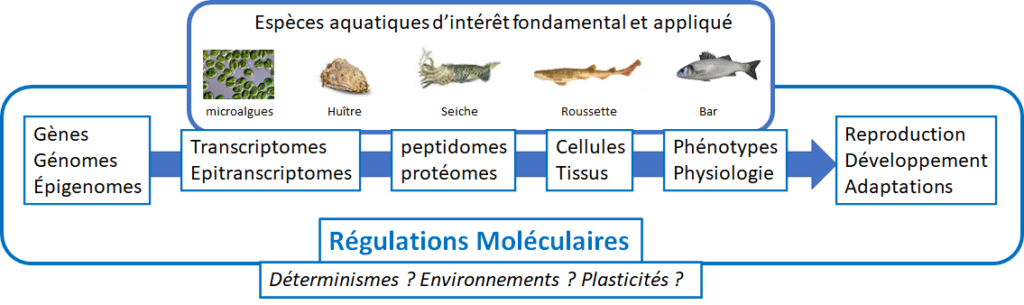
The studied species stand at key positions across a broad phylogenetic range and include both wild and aquaculture-targeted organisms. These models are of both fundamental and applied interest, and include:
- Microalgae (aquaculture-relevant phyla)
- Bivalve mollusks: Crassostrea (e.g. Magallana) gigas (Pacific oyster), Ruditapes philippinarum (Manila clam)
- Gastropods: Buccinum undatum (whelk)
- Cephalopods: Sepia officinalis (common cuttlefish)
- Cartilaginous fish: Scyliorhinus canicula (small-spotted catshark)
- Teleost fish: Dicentrarchus labrax (European seabass), Heterotis niloticus (African bonytongue, a Cameroonian species)
The team aims to understand the physiological specificities of these organisms and how they respond to environmental stimuli.
Areas of research
Axis 1 : Molecular and cellular determinants of key control systems of the biological cycle
This axis deciphers the molecular and cellular pathways involved in the fate of cells engaged in, or responsible for, critical physiological processes. Emphasis is placed on reproductive and developmental physiology, including the study of factors and mechanisms that govern the balance between the stem cell identity and the differentiation of germline and somatic lineages. At the molecular level, gene expression regulation, particularly via epigenetic mechanisms (nucleic acid modifications, chromatin remodeling), is a central focus.
Research also investigates neuroendocrine, immune, and nutritional control pathways during key life cycle stages, through the characterization of communication networks (neuroendocrinology), bioactive molecules, receptor deorphanization, and the molecular mechanisms of sex determination.
Axis 2: Adaptive and evolutionary consequences of the plasticity of physiological and biological cycle control systems
Physiological regulatory processes play a crucial role in organism adaptation to environmental conditions. While these controls are fundamentally genetic, they are also modulated by environmental factors. This axis focuses on the plasticity of these systems in response to climate change, human activity, and contamination.
The adaptive and evolutionary implications of this plasticity are particularly relevant for the sustainable management of fisheries and aquaculture species, considering both immediate impacts (e.g., rearing conditions, immunity, spawning) and long-term effects, including intergenerational and transgenerational impacts (e.g., epigenetic and epitranscriptomic inheritance, predictive modeling).
Expertise and Methodologies
Team members have strong expertise in complementary approaches, including:
- Chromatin, genetic, and epigenetic studies: qPCR, NGS and Nanopore sequencing, bioinformatics, immunoprecipitation
- Transcriptomics, proteomics, and peptidomics: NanoLC-MS/MS, reverse endocrinology
- Cellular and histological analyses: cell culture and sorting, qualitative and quantitative microscopy, histology
- Molecular characterization: immunolabeling, in situ hybridization
- Functional tests (in vivo and in vitro): exposure studies, microinjections, transplantations, gene editing
These tools are applied in controlled laboratory experiments, field studies, and modeling approaches (e.g., DEB models), to interpret results within a broader ecological and evolutionary context.
Recent publications
Journal articles
2026
- ref_biblio
- Sauger, C., Quinquis, J., Kellner, K., Berthelin, C., Lepoittevin, M., Elie, N., & Dubroca, L. (2026, Feb). Comparing Macroscopic and Quantitative Histological Methods to Determine Sexual Maturity in the Female European Plaice. Animals, 16 (519), 1-18. <10.3390/ani16030519>. <hal-05497774>.
2025
- ref_biblio
- Bouras, H., Quesnelle, Y., Trancart, S., Goux, D., Blin, J.-L., Savary, M., Houssin, M., & Zatylny-gaudin, C. (2025, Nov). Phenotypical and Genomic Characterization of the Mollusk Pathogen Francisella halioticida. MicrobiologyOpen, 14 (6), e70172. <10.1002/mbo3.70172>. <hal-05388653>.

- ref_biblio
- Clairet, N., Auger, H., Arredondo-espinoza, R.-C., Koechlin, H., Bernay, B., Manoury, L., Goux, D., & Rivière, G. (2025, Nov). m\textlesssup\textgreater6\textless/sup\textgreaterA-RNA epitranscriptomes regulate splicing and neuronal development in the Pacific oyster Crassostrea gigas. Genomics, 117 (6), 111142. <10.1016/j.ygeno.2025.111142>. <hal-05349821>.

- ref_biblio
- Jeanne, F., Pilet, S., Combarnous, Y., Bernay, B., Dufour, S., Favrel, P., & Sourdaine, P. (2025, Jul). Pleiotropic signaling of single-chain thyrostimulin (GPB5-GPA2) on homologous glycoprotein hormone receptors (ScFSHR, ScLHR, ScTSHR) in the elasmobranch Scyliorhinus canicula reproduction. Molecular and Cellular Endocrinology, 604, 112553. <10.1016/j.mce.2025.112553>. <hal-05044952>.

Conference papers
2025
- ref_biblio
- Bouras, H., Corre, E., Bernay, B., Blin, J.L., Savary, M., Houssin, M., & Zatylny-gaudin, C. (2025, Sep). Immune response of Mytilus edulis challenged with low-virulence and high-virulence isolates of F. halioticida. Paper presented at the 22nd International Conference on Disease of Fish and Shellfish, Heraklion - Crète, Greece. <hal-05240201>.
- ref_biblio
- Braquart-varnier, C., Dussert, Y., Moumen, B., Pailler, L., Peccoud, J., Verdon, J., & Zatylny-gaudin, C. (2025, May). Identification of new antimicrobial peptides in Armadillidium vulgare using PepTraq.. Paper presented at the REID 2025, Lyon, France. <hal-05518405>.
- ref_biblio
- Clairet, N., Auger, H., Arredondo-espinoza, R.-C., Koechlin, H., Bernay, B., Goux, D., & Rivière, G. (2025, May). M6A-RNA EPITRANSCRIPTOMES CONTROL NEURONAL DEVELOPMENT AND GENE SPLICING IN THE OYSTER. Paper presented at the EPIMAR2025 - Third International Symposium on Epigenetics in Marine and Aquatic Research, Barcelona, Spain. <hal-05349827>.
- ref_biblio
- Clairet, N., Auger, H., Bouillet, A., Roberto Carlos, A.-E., Kellner, K., Elie, N., Nadège, V.-N., & Rivière, G. (2025, May). PUTATIVE M6A-DEPENDENT DOSAGE COMPENSATION IN TRIPLOID OYSTERS SUGGESTS EXTENSION OF THE X CHROMOSOME INACTIVATION PARADIGM OUTSIDE MAMMALS. Paper presented at the EPIMAR2025 - Third International Symposium on Epigenetics in Marine and Aquatic Research, Barcelone, Spain. <hal-05549463>.

- ref_biblio
- Pred'homme, M., Zatylny-gaudin, C., Villain-naud, N., Dubos, M.-P., Goux, D., Elie, N., & Henry, J. (2025, May). Post-reproductive Neurodegeneration in Cuttlefish: Unveilling an Alzheimer’s-like Tauopathy?. Paper presented at the Meeting of the Club of Invertebrate Neurobiology, Lyon, France. <hal-05548920>.

- ref_biblio
- Pred'homme, M., Zatylny-gaudin, C., Villain-naud, N., Dubos, M.-P., Goux, D., Elie, N., Bernay, B., Corre, E., & Henry, J. (2025, Jun). The cuttlefish $Sepia\ officinalis$: a relevant invertebrate model for the study of Alzheimer's disease?. Paper presented at the FAAST 2025 - France Alzheimer Actualités Scientifiques et Thérapeutiques, Paris, France. <hal-05548904>.
















